Discovering unique microbes made easy with new software platform
Microbes are foundational for daily life on Earth. These tiny organisms play a big function in all the things from reworking sunlight into the basic molecules of lifestyle. They aid to produce considerably of the oxygen in our ambiance. They even cycle nutrients among air and soil. Experts are continually locating interactions in between microbes and vegetation, animals, and other macroscopic lifeforms.
As genomic sequencing has advanced, researchers can look into not only isolated microbes, but also total communities of microorganisms—known as microbiomes—based on DNA identified in an ecosystem. The genomes extracted from these communities (metagenomic sequences) can discover the organisms that carry out biogeochemical procedures, contribute to health and fitness or disorder in human gastrointestinal microbiomes, or interact with plant roots in the rhizosphere.
The Section of Power Programs Biology Knowledgebase (KBase) lately produced a suite of attributes and a protocol for doing refined microbiome assessment that can speed up exploration in microbial ecology.
The popular adoption of DNA sequencing in microbiology has generated big amounts of genomic info. Researchers have to have computational instruments to recuperate higher-top quality genomes from environmental samples to realize which organisms reside in an ecosystem and how they could possibly interact. The mix of usability, info, and bioinformatics resources in a community on the web useful resource would make KBase a uniquely highly effective world-wide-web platform for executing this endeavor.
These new options in KBase will let biologists to get genomes from microbiome sequences with uncomplicated-to-use program run by Division of Power computational means. This will lower the time expected to system sequencing info and characterize genomes. Researchers can use KBase to collaboratively review genomics data and establish study communities to solve common complications in microbial ecology.
Obtaining genomes of uncultivated microbes instantly from the ecosystem using DNA sequencing is a latest progress that will allow scientists to discover and characterize novel organisms. Sequencing the DNA of all the microbes in a offered ecosystem produces a “metagenome.” Accomplishing genetic investigation of metagenomes has emerged as a way to check out microbial traits and behaviors and local community interactions in an environmental context.
Techniques for getting metagenome-assembled genomes (MAGs) have different levels of achievement, relying on the strategies employed. An increasing amount of researchers crank out microbiome sequences, but numerous do not have all set accessibility to the expertise, resources, and computational assets important to extract, evaluate, and assess their genomes.
The KBase group extra and current a number of metagenome analysis equipment, data kinds, and execution capabilities to offer scientists the equipment that accelerate the discovery of microbial genomes and uncover the genetic probable of microbial communities.
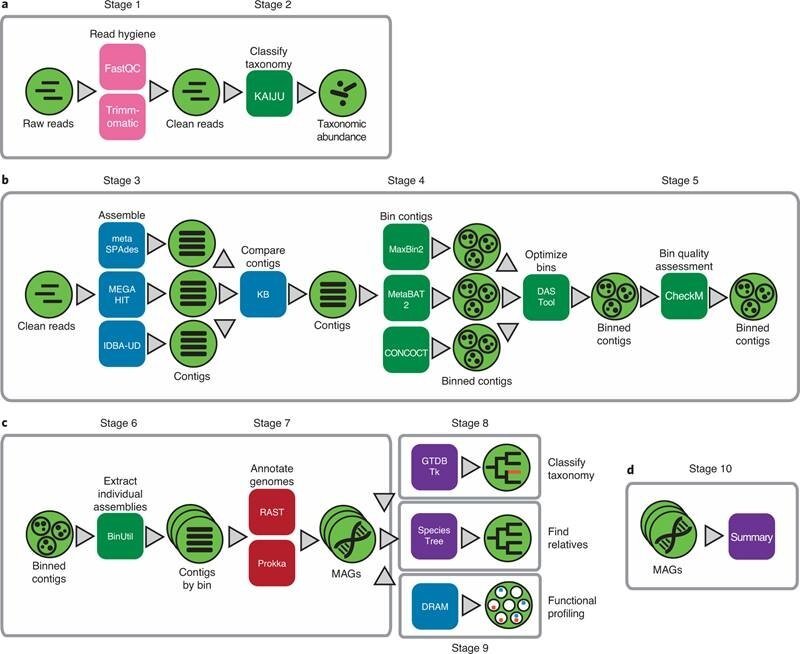
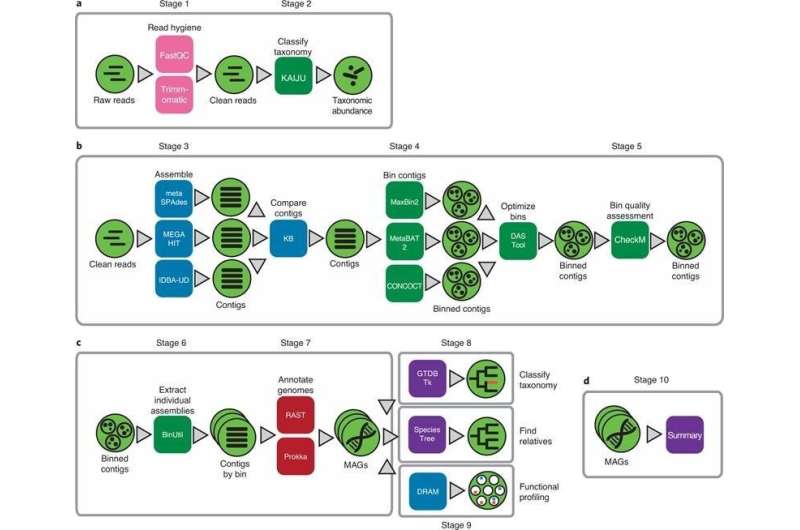
Their recent paper in Character Protocols offers a collection of analysis actions, utilizing KBase applications and facts products and solutions for extracting higher excellent MAGs from metagenomes. These abilities, like computing, details storage, and sharing of info and analyses, are delivered totally free to the general public by way of the KBase web system. This protocol will allow researchers to equally crank out putative genomes from organisms only located in the setting and assess them with tools to recognize who they are, what they are carrying out, who they are interacting with, and their part in the ecosystem.
Extra info:
Dylan Chivian et al, Metagenome-assembled genome extraction and examination from microbiomes working with KBase, Character Protocols (2022). DOI: 10.1038/s41596-022-00747-x
Quotation:
Finding distinctive microbes produced uncomplicated with new computer software system (2023, January 30)
retrieved 2 February 2023
from https://phys.org/news/2023-01-special-microbes-effortless-software package-platform.html
This document is matter to copyright. Apart from any reasonable dealing for the purpose of personal study or investigation, no
portion could be reproduced without the need of the penned permission. The articles is furnished for info needs only.
