New Technology Uncovers Hidden Mitochondrial DNA Mutations
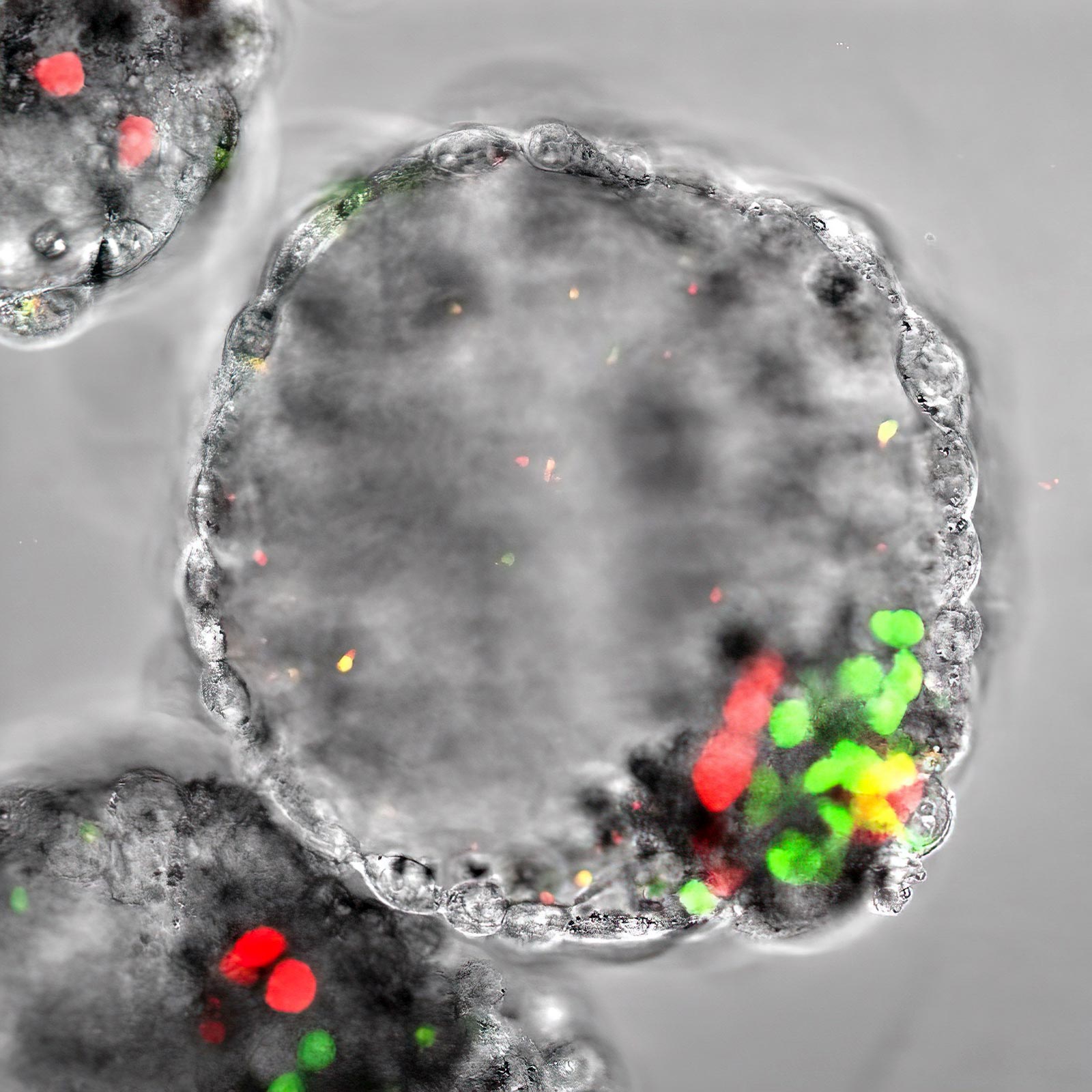
A human blastocyst-like artificial embryo known as blastoid showing the presence of an enveloping layer of added-embryonic cells, a blastocoel-like cavity, epiblast cells (green, offering rise to the foreseeable future embryo) and hypoblast cells (crimson, giving rise to the foreseeable future amnion). iMiGSeq was employed to sequence mtDNA in a one blastoid to design the dynamics of mtDNA mutations for the duration of human embryogenesis. Credit score: © 2023 KAUST Mo Li
A substantial-throughput single-mobile single-mitochondrial genome sequencing know-how recognized as iMiGseq has offered new insights into mutations of mitochondrial
An international team of researchers, led by KAUST stem cell biologist Mo Li, has now quantitatively depicted the genetic maps of mtDNA in single human oocytes (immature eggs) and blastoids (stem cell-based synthetic embryos).[1] This has uncovered molecular options of rare mtDNA mutations that bring about maternally inherited disorders.
Mitochondria, the “powerhouses” of cells, enjoy a important purpose in cellular conversation and metabolic rate. Human mtDNA is a circular genome that contains 37 genes, encoding 13 proteins and a noncoding D-loop location. Heteroplasmic mutations, inherited from egg cells, can bring about congenital disorders, like maternally inherited Leigh syndrome, and are associated with late-onset intricate health conditions.
“Next-era sequencing has been applied to sequence mtDNA and implicated heteroplasmic mutations as considerable contributors to metabolic ailment. Still the knowledge of mtDNA mutations continues to be minimal thanks to the constraints of standard sequencing systems,” claims lead writer Chongwei Bi.
“Our new iMiGseq system is major due to the fact it allows comprehensive sequencing of unique mtDNA in one cells, enabling for unbiased, significant-throughput base-resolution evaluation of whole-duration mtDNA,” states Bi. iMiGseq resolves various crucial issues in the industry.
Employing third-technology nanopore sequencing engineering, the researchers have characterised mtDNA heteroplasmy in one cells and described the genetic functions of mtDNA in single oocytes. They have examined mtDNA in induced pluripotent stem cells derived from sufferers with Leigh syndrome or neuropathy, ataxia or retinitis pigmentosa (NARP). This has revealed intricate patterns of pathogenic mtDNA mutations, such as solitary nucleotide variants and substantial structural variants. “We ended up ready to detect rare mutations with frequencies far under the traditional detection threshold of a person per cent,” claims Mo Li.
In an additional experiment working with the new technologies, iMiGseq revealed the potential risks of surprising significant raises in the frequency of off-concentrate on mutations, known as heteroplasmy, in a mitochondrial genome editing approach named mitoTALEN – a genome editing tool that cuts a precise sequence in mitochondrial DNA. It is utilized to slash a mutation that causes mitochondrial encephalomyopathy and stroke-like episodes syndrome in affected person-derived induced pluripotent stem cells.
“This highlights the strengths of whole-size mtDNA haplotype examination for being familiar with mitochondrial DNA heteroplasmy improve other distant mtDNA genetic variants could be unintentionally afflicted by the editing of a genetically connected ailment-relevant mutation and there is a need for ultrasensitive strategies to evaluate the protection of enhancing approaches,” states Li.
The researchers also used iMiGseq to analyze single human oocytes from wholesome donors and one human blastoids, artificial embryos produced from stem cells, to establish scarce mutations undetectable with typical next-generation sequencing. These reduced-amount heteroplasmic mutations, potentially inherited by means of the female germline, are linked to mitochondrial health conditions and most cancers.[2]
The iMiGseq strategy supplies a novel suggests to precisely depict the complete haplotypes of particular person mtDNA in solitary cells, providing an excellent platform for describing the bring about of mitochondrial mutation-connected conditions, analyzing the protection of several mtDNA enhancing techniques and unraveling the inbound links in between mtDNA mutations, ageing and the development of elaborate conditions.
References:
- “Quantitative haplotype-resolved analysis of mitochondrial DNA heteroplasmy in Human one oocytes, blastoids, and pluripotent stem cells” by Chongwei Bi, Lin Wang, Yong Lover, Baolei Yuan, Samhan Alsolami, Yingzi Zhang, Pu-Yao Zhang, Yanyi Huang, Yang Yu, Juan Carlos Izpisua Belmonte and Mo Li, 4 April 2023, Nucleic Acids Investigation.
DOI: 10.1093/nar/gkad209 - “Single-cell specific total-size mtDNA sequencing by iMiGseq uncovers surprising heteroplasmy shifts in mtDNA editing” by Chongwei Bi, Lin Wang, Yong Enthusiast, Baolei Yuan, Gerardo Ramos-Mandujano, Yingzi Zhang, Samhan Alsolami, Xuan Zhou, Jincheng Wang, Yanjiao Shao, Pradeep Reddy, Pu-Yao Zhang, Yanyi Huang, Yang Yu, Juan Carlos Izpisua Belmonte and Mo Li, 31 March 2023, Nucleic Acids Study.
DOI: 10.1093/nar/gkad208
