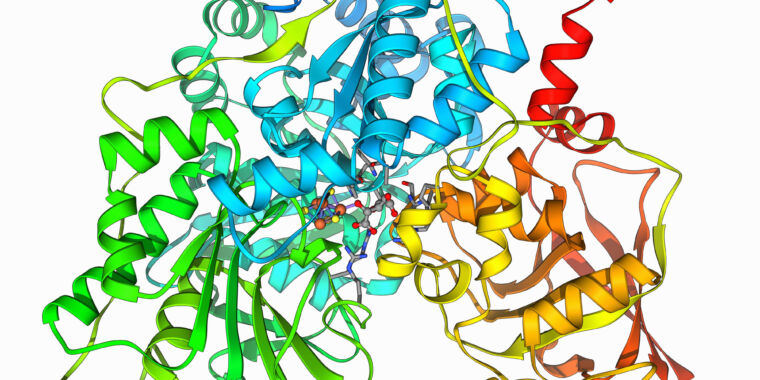
Have a protein you want inhibited? New software can design a blocker
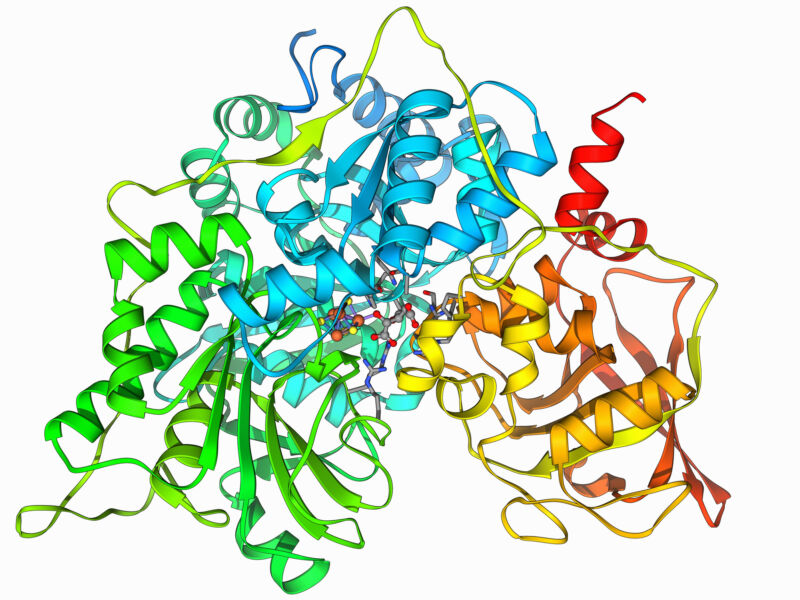
Thanks in part to the substantial assortment of shapes they can undertake and the chemical environments people styles build, proteins can carry out an remarkable number of features. But there are lots of proteins we want did not operate rather so effectively, like the proteins on the surfaces of viruses that let them latch on to new cells or the ruined proteins that induce cancer cells to develop uncontrollably.
Ideally, we would like to block the important web pages on these proteins, restricting their potential to do hurt. We’ve noticed some progress in this spot with the introduction of a selection of smaller-molecule drugs, such as just one that seems effective versus COVID-19. But that kind of drug enhancement normally final results in substances that, for one purpose or an additional, really don’t make powerful medication.
Now, researchers declared they have made computer software that can design and style a separate protein that will adhere to a goal protein and likely block its activity. The software package was diligently developed to limit the processing calls for of a computationally intricate method, and the full factor advantages from our skill to do massive-scale validation checks working with molecular biology.
How do you work out that?
Proteins are a string of amino acids, with the cell making use of 20 various amino acids that have some distinct chemical attributes. Interactions amid these amino acids allow for the proteins to undertake a three-dimensional configuration, which frequently provides key amino acids jointly in a person area to kind an “active web-site” that is significant for the protein’s operate. Many medicines are intended to match into this lively web site, which can retain a protein from operating.
An option is to protect the web site up with a little something massive that sticks to it. And just one of the least complicated items to get to stick to a protein is one more protein. This solution has some limitations when it comes to drug development. But the biggest challenge is coming up with a protein that specially sticks to another. Given that every single website link in a protein’s chain can be occupied by a single of 20 diverse amino acids, the amount of attainable proteins grows huge right after the chain is just a several backlinks long. So exploring all feasible interacting proteins is absolutely intractable as a computation.
The new operate, which was completed by a substantial US-dependent collaboration, targeted on minimizing the computational complexity. For the new approach, the researchers resolved to concentrate on manufacturing tons of weak interactions alternatively than exclusively designing a couple of sturdy ones. So the process begun by having each individual of the 20 personal amino acids and screening the toughness of its interactions at billions of different sites on the area of the goal protein. The specifics of these interactions were being then stored for later on use.
Individually, the researchers developed up a databases of about 35,000 limited proteins (about 50-65 amino acids lengthy) that fashioned a steady main. These proteins have been then put up coming to the concentrate on protein. The algorithm would then ascertain where each and every amino acid was in near proximity to the target and then use the previously outcome to pick which amino acid would have the strongest interactions at that spot.
This process still left a a great deal smaller sized range of protein interactions that necessary to be screened in element. The scientists utilized a “packing” algorithm to see how the two proteins match together and then approximated the toughness of their binding.
